Computational Biology Development
Advertisement
JigCell for Mac OS X v.7.1.0
JigCell is an effort to develop and maintain an open source modeling environment for computational biology.
Advertisement
JigCell for Windows v.7.1.0
JigCell is an effort to develop and maintain an open source modeling environment for computational biology.
Electronic Cancer System Studio v.1.0.0.1
A computational model development and simulation environment for cancer systems biology.
Pathway Tools v.15.0
Pathway Tools is a comprehensive symbolic systems biology software system that supports several use cases in bioinformatics and systems biology: - Development of organism-specific databases - Scientific Visualization, web publishing - Visual anal
JigCell v.7. 1. 2000
JigCell is an effort to develop and maintain an open source modeling environment for computational biology.
Insilicos Cloud Army v.2.0.0
Platform for parallel computation in the Amazon cloud, including machine learning ensembles written in R for computational biology and other areas of scientific research.
NeoBio v.1.0
NeoBio is a Java class library of Computational Biology Algorithms.
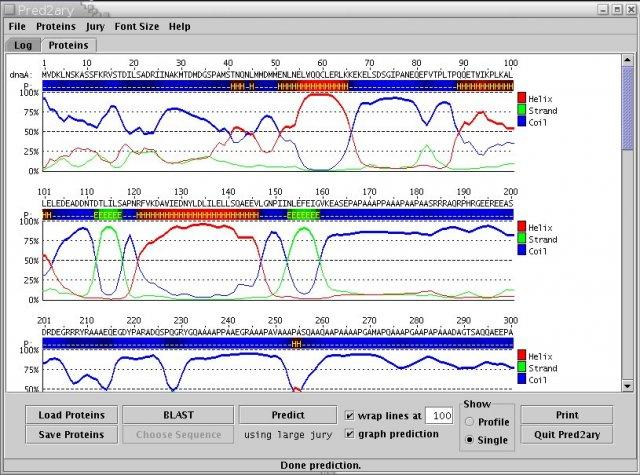
StrBio java class libraries v.b.1.2
Java class libraries for structural biology development: includes protein format conversion tool, printf-based text formatting, Pred2ary secondary structure prediction, neural net library, Hooke-Jeeves global optimizer, and misc.
Strainer v.Release 23
View metagenomics data with this tool. Strainer is a multi-platform visualization tool for metagenomics data. It allows the user to easily browse the variation present in shotgun sequence data from non-clonal samples.
AxonTracker v.1.0
Track axon structures with the help of this tool.
Genome Explorer v.1.0
Search sequence data with this tool. Genome Explorer offer users a single interface for some useful bioinformatics tools. It provides additional functionality for searching sequence data (for particular groups of nucleotides or amino acids) and